The mission of the Academy's Institute for Biodiversity Science and Sustainability (IBSS) is to gather new knowledge about life's diversity and the process of evolution—and to rapidly apply that understanding to our efforts to regenerate life on Earth.
Center for Comparative Genomics
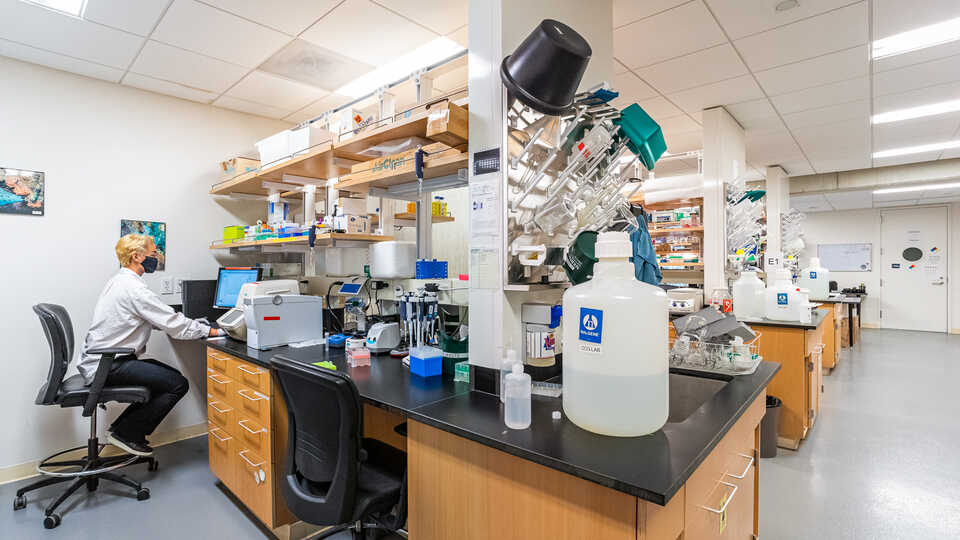
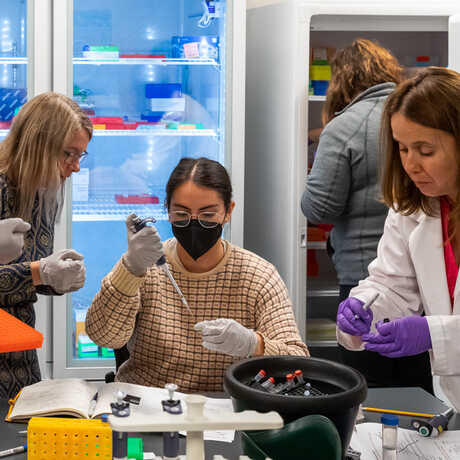
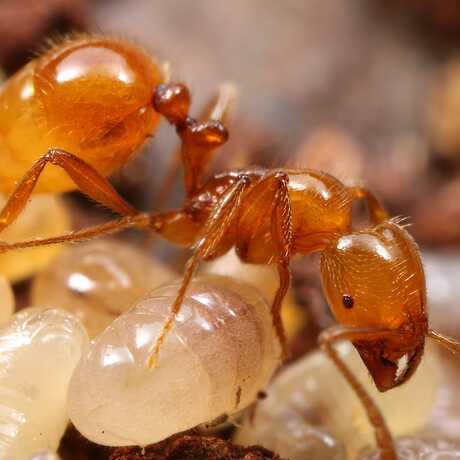
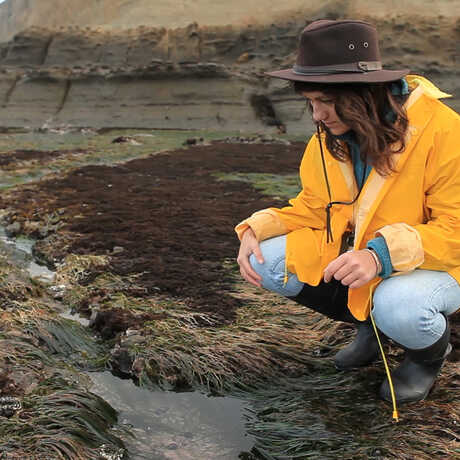
The Center for Comparative Genomics (CCG) serves as the Academy's central hub for genomics and sequencing. Our facilities include a fully-equipped genomics laboratory, a frozen DNA CryoCollection, and high-performance computing resources. The CCG focuses on museum genomics, employing the latest molecular and computational techniques and utilizing natural history specimens to explore questions relevant to biodiversity, conservation, and regeneration. Our primary mission is to spearhead groundbreaking research in museum genomics at the Academy. The CCG engages in diverse projects spanning the breadth of the tree of life and spanning the globe—From the Caribbean coral reefscapes and Galápagos birds to frogs of Cameroon, California native plants, and bioluminescent fish microbiomes in the Indo-Pacific. In addition to our research efforts, we are dedicated to training and supporting the next generation of biodiversity researchers. As leaders in museum genomics, the CCG hosts workshops and training sessions for local researchers interested in the field. Our aim is to create a supportive environment for scientific exploration and knowledge sharing.
More about the CCG
More about CCG Projects